G4X™ Spatial Research Challenge
Apply for one of two in situ multiomic research award tracks on G4X: a funded 1,000-sample cohort study or a 3D reconstruction study using serial FFPE sections.
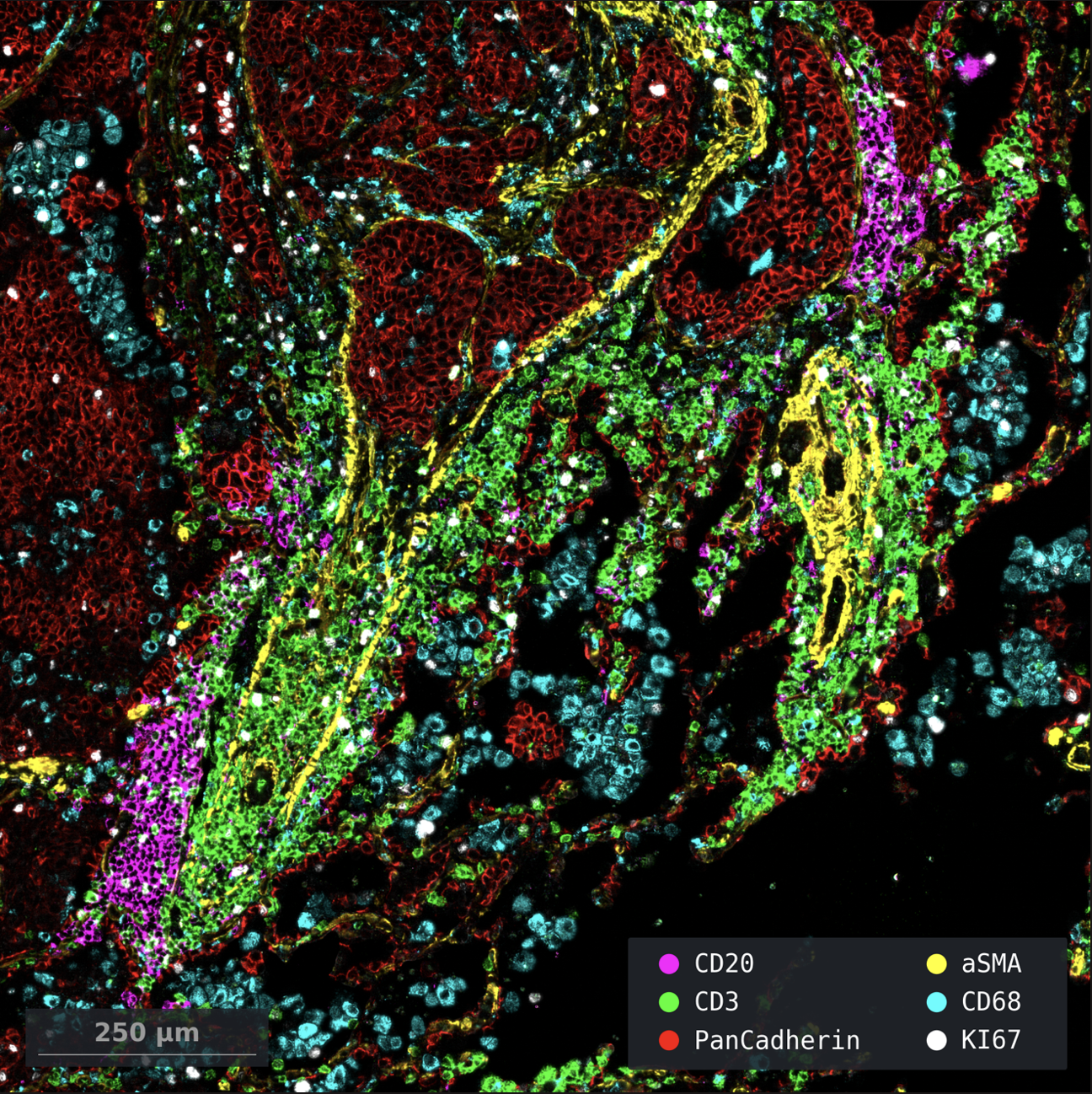
Both tracks include up to 1,000-plex RNA, 16-plex protein, and fH&E in the same FFPE section.
One Challenge. Two Ways to Push Spatial Further.
The G4X Spatial Research Challenge is designed to support ambitious studies that take advantage of multimodal spatial analysis on FFPE tissue.
For researchers pursuing cohort-scale discovery, the flagship award supports a fully funded 1,000-sample study. For researchers focused on tissue architecture and three-dimensional biology, the second award track supports 3D reconstruction from serial FFPE sections.
Together, these two tracks are intended to support a broader range of high-impact studies, from large translational cohorts to spatially rich 3D tissue investigations.
1,000 Sample Spatial Study
A single winning proposal will be awarded a funded 1,000-sample multimodal spatial study run by Singular Genomics on G4X. Runner ups will be considered for additional awards.
Included in every sample
- up to 1,000-plex RNA*
- 16-plex protein
- fH&E
Designed for
- translational biomarker studies
- therapy response and resistance
- large retrospective cohorts
- AI-ready spatial dataset generation
Best for
Studies where cohort size, statistical power, and multimodal tissue insight are central to the scientific question.
Details
Sample size is either 4.5x4.5mm or 10x10mm or a mix, 1,000 samples relates to smaller sample size, max number of larger samples is 320 (for a total of 320cm2 imaging space on 32 G4X flow cells)

Courtesy of Dr. Shanye Yin, Assistant Professor in the Department of Pathology at Albert Einstein College of Medicine
3D Spatial Study
A single winning proposal will receive a funded 3D in situ multiomic study run by Singular on G4X.
Option A
- up to 8 samples
- 40 µm reconstruction depth per sample
- 4.5 × 4.5 mm area
Option B
- Up to 4 samples
- 80 µm reconstruction depth per sample
- 4.5 × 4.5 mm area
Option C
- Up to 2 samples
- 50 µm reconstruction depth per sample
- 10 × 10 mm area
Included in every sample
- Up to 1,000-plex RNA
- 16-plex protein
- fH&E
Best for
Studies where tissue architecture, cell-cell relationships, and three-dimensional structure are central to the biological question.

About G4X™
- ~10 Times More Throughput - Process up to 128 samples per run with 40cm² of flexible imaging area.
- Breakthrough Pricing - Spatial from $240 per sample.
- Unified Multiomics - 500-plex RNA, 18 proteins, and fH&E from a single FFPE section.
- Direct-Seq™ - In situ sequencing provides novel genomic information in spatial context

Continue Exploring
View the Poster